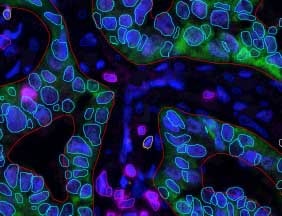
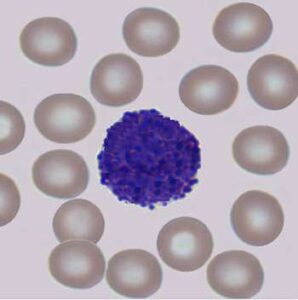
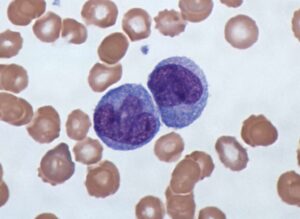
Studying the behavior of cells in-vitro is one of the most fundamental research tools in biology. Studies conducted under the microscope improve our understanding of cell-cell interaction, motility, and reaction to different biochemical conditions. In addition, cell counting and sorting, which are mainly done manually, are routine laboratory tasks conducted under the microscope.
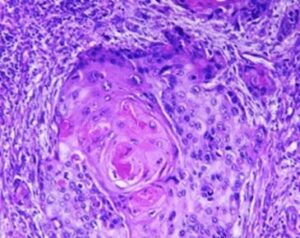
Manual inspection is limited to tracking a relatively small number of cells in short periods of time and it is prone to human error. On the other hand, computer vision algorithm can be utilized to perform fast scanning, segmentation and tracking of large cell population over long periods of time. Moreover, tailor-made algorithm can be put to quickly scan, analyze and output results, e.g. alert on histological abnormalities, saving thus labor and time, hence enabling a more timely therapy.
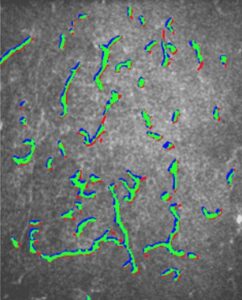
Whichever the end goal of the cells tracking research, when cells are the object of interest, the primary objective is segmenting out the cells in the image. A successful segmentation of cells allows then a more in-depth analysis of their morphology, contact, position over time and behavior.
At this stage of the cells tracking we can already give quantitative information about the cell colony: number of cells, size distribution, number of contacts and their position. Further characterization of each segmented region, based on combination of morphological and color features, permits further extraction of valuable information.
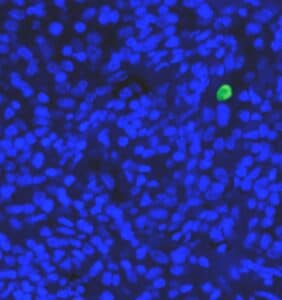
Essays dealing with cell production of proteins of interest rely heavily on the expression level to characterize response. Once a colony has been segmented, the fraction of cells expressing a certain protein of interest can be given. Moreover, the expression level and the correlation to position in the colony can be estimated.
A per cell extraction of features permits tracking cells throughout acquisition cycle by correlating features in two subsequent time points. Along with the integration of spatial and morphological information, position and condition of the cell can be given at any moment in time.
Automatic segmentation and tracking of cells is a non-trivial task. It requires the ensemble of knowledge for methods such as segmentation, microscopy, tracking deformable objects and machine learning. Our team at RSIP Vision has gained expertise in these fields enabling us to develop and deliver cutting-edge solutions for over 2 decades.


 Microscopy
Microscopy