This article was first published on Computer Vision News of March 2022.
Mara Graziani recently completed her Ph.D. at the University of Geneva and the University of Applied Sciences of Western Switzerland (Hes-so Valais). She researches interpretability techniques for deep learning models used in healthcare. She will now embrace a new postdoctoral adventure at IBM Research Zurich and the Zurich University of Applied Sciences (ZHAW) trying to apply interpretability on systems biology. Congrats, Doctor Mara!
The models that we work with in machine learning are approximations based on assumptions that are never exactly true. Even if wrong, some models may be useful.
I find it rather compelling to understand when a model is useful for the clinical application. In this field, mistakes come at a high cost, and understanding the limitations of a model is fundamental. If pitfalls are uncovered, new models can be built to be more reliable and trustworthy than the already existing ones.
Being the end-users of our developments, physicians should be part of the model evaluation process, though they often ignore the processes of feature extraction, model selection and training. If they were explained which features are used by the model to obtain a certain output, they would then be able to predict eventual failures and unexpected outcomes.
So can we make Deep Learning models for medical image classification more understandable to physicians? How can this analysis be used to improve model generalization?
Physician-friendly Explanations
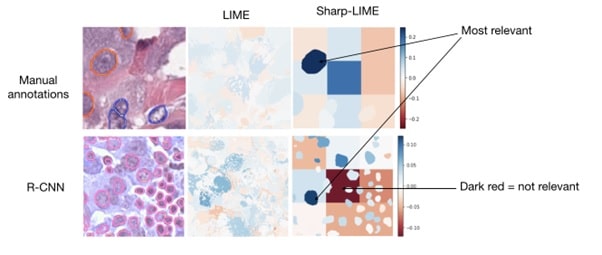
We proposed visual and concept-based explanations that are understandable by pathologists, focusing on the application of breast cancer detection in digital scans of tissue samples. Our work on Sharp-LIME uses nuclei contours to generate neat heatmaps that highlight the relevance of individual nuclei to the prediction. With Regression Concept Vectors I have generated explanations in terms of nuclei morphology and appearance.
These works improved the understandability of the explanations by physicians and generated new insights on the CNN mechanics for cancer detection.
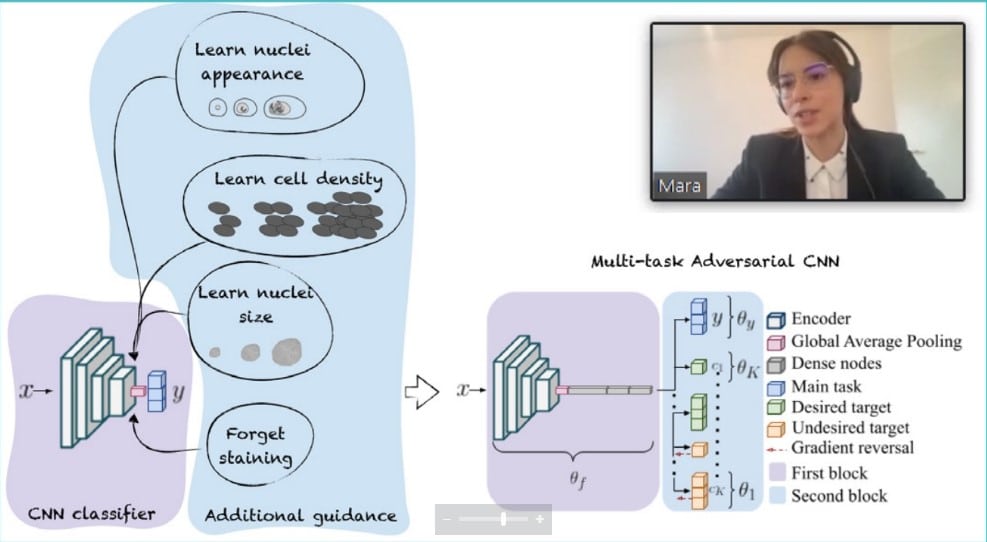
CNNs were shown to be biased towards earning texture differences rather than shapes and scale. These biases may affect the generalization if new scanning protocols or equipment are adopted by the laboratory. To counter this limitation, we developed a multi-task adversarial architecture that encourages the learning of clinical features. This model has an additional branch for each clinical feature that is important to predict tumorous regions, e.g. nuclei density, size and appearance. The shift across data from multiple institutions is handled by an additional adversarial branch that removes the domain information from the internal features. This architecture outperforms conventional CNN-based approaches for the prediction of tumor vs. normal regions in histology slides.
Acknowledgement
This research was supported by the EU Horizon 2020 projects PROCESS, AI4Media and ExaMode.
Keep reading the Medical Imaging News section of our magazine.
Read about RSIP Vision’s R&D work on Medical Image Analysis.