This article was first published on Computer Vision News of December 2021.
Chen (Cherise) Chen recently completed her Ph.D. at Imperial College London (ICL). Her main research interest lies in machine learning and computer vision, and their use in medical image analysis, with the ultimate goal of improving machine learning algorithms’ robustness and reliability in realworld healthcare applications. She will continue to work as a postdoctoral researcher in her current group: BioMeDIA. Chen would like to thank her supervisors Daniel Rueckert and Wenjia Bai for their continued support and guidance during this long and exciting journey. Congrats, Doctor Chen!
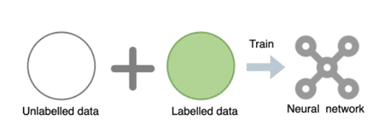
Deep neural networks have become a powerful tool to process and analyze medical images, with great potential to accelerate clinical workflows and facilitate largescale studies. However, in order to achieve clinically acceptable performance at deployment, these networks generally require massive labeled data collected from various domains (e.g., hospitals, scanners). Such large labeled datasets are rarely available in practice, due to the high collection and labeling costs as well as data privacy issues. So, how can we improve the domain generalization and robustness of neural networks for medical imaging using limited labeled data?
Learning with unlabelled data
We consider exploiting the value of unlabelled data, which is cheap and often widely available. We develop a novel adversarial data augmentation (DA) method with realistic image transformations (adv bias, adv chain) to augment labeled and unlabelled images for more effective consistency regularization. This adversarial DA can be applied as a plug-in module in both supervised and semi supervised medical image segmentation frameworks.
We further propose an unsupervised style transfer method with a cascaded U-net for unsupervised domain adaptation, where there is a distributional gap between source domain images (labeled) and target domain images (unlabelled). We first train an imagestyle translation network by utilizing a set of labeled bSSFP MR images (source domain) and unlabelled LGE MR images (target domain). We then employ the trained network to translate labeled bSSFP images to be LGE-like images, which are then used to train an LGE segmentation network. Our approach does not require labeled LGE images for training, yet it can surprisingly achieve high segmentation accuracy on LGE images, ranking 1st in the public multi-sequence cardiac MR segmentation challenge [More].
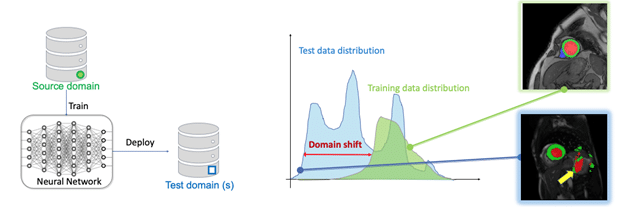
Learning from limited, single-domain data for improved cross-domain performance
In the worst scenario where we only have access to limited labeled data from a single domain for training, the model can easily fail to predict images from unseen test sets due to the domain shift problem (see Figure above). A natural solution is data augmentation, which simulates the domain shifts at training. We first design a general training and testing pipeline with data normalization and augmentation to improve cardiac image segmentation performance across various unseen domains. We further propose a latent space data augmentation method with a cooperative training framework to enhance model robustness against unseen domain shifts and imaging artifacts. By performing random/ targeted code masking in the latent space, a set of challenging examples are generated to strengthen the network capacity. This latent-space masking-based data augmentation method is effective and practical, as it does not require domain knowledge and expertise to design a particular image augmentation/corruption function.
Keep reading the Medical Imaging News section of our magazine.
Read about RSIP Vision’s R&D work on Medical Image Analysis.