Using Deep Neural Networks for Vessel Segmentation in Fundus Images
Vessel segmentation methods based on image processing techniques have long been utilized to delineate the vascular tree in clinical imaging. Many vessel segmentation algorithms exist and have been widely demonstrated in ophthalmology, where noninvasive modalities enables the external inspection of the condition and structure of blood vessels. With limited rates of success, the segmentation methods have been replaced by more sophisticated methods for detecting vascular tree based on learning networks, and more recently developments in Deep Neural Networks (DNN). This article explores vessel segmentation using deep learning, including DNN and CNN.
The detection of arteries in ophthalmology is a basic procedure for the diagnosis of retinal pathologies. Retinal pathologies are commonly seen and are indicative of patients with diabetes and hypertension. Early detection and accurate identification of the pathological progression is essential for preventing further deterioration, and in severe cases, vision loss.
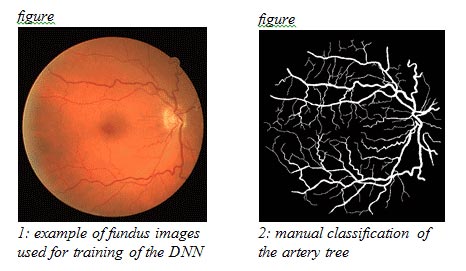
Pathologic progression is oftentimes characterized by abnormal an vascular arrangement in the eye. Significant prognostic factors include geometric characteristics of the vessels – such as diameter, length and branching angles – as well as the distinction between arteries and veins, and their mutual arrangements in the eye. Those features are visible by fundus imaging (see example in figure 1), using a contrast agent to enhance the contrast of the vessels against their background. While manual examination of the progression of the disease is time consuming, ofttimes requiring integration and comparison of information from several modalities, algorithmic means emerge as a time-saving, fast and accurate means of detecting and tracking pathologies characterized by abnormalities in the vascular tree.
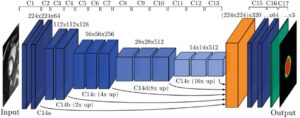
Segmentation of the vascular tree forms the first step towards the construction of an algorithmic basis for pathological detection. For this end, several binary classifiers have been proposed in the literature. One of the most recent, by Melinscak et al. of the University of Zagreb, offers a DNN approach for the construction of the binary classifier. In essence, for each pixel in a given image, the classifier relates it to a class – a vessel or not a vessel pixel (see the example of manual segmentation in figure 2, used for training of DNN). The choice of DNN seems a natural approach in this context, for which heterogeneity in the roles of processing layers in the network is required for recognizing different geometrical and spatial features of vessels, e.g. orientation of edges vs. color features. Melinscak’s 10 layered DNN underwent training in some of its layers, which included the filtering of the image between layers and the data propagated between them. It was then put to the test. By measuring the area under the Receiver Operating Characteristic (ROC), the researchers report an average accuracy of 0.96 to 0.98 in several experiments, which demonstrates the high potential and usability of DNN for automatic vessel segmentation.
The detection of arteries in ophthalmology is a basic procedure for the diagnosis of retinal pathologies. Retinal pathologies are commonly seen and are indicative of patients with diabetes and hypertension. Early detection and accurate identification of the pathological progression is essential for preventing further deterioration, and in severe cases, vision loss.
Pathologic progression is oftentimes characterized by abnormal an vascular arrangement in the eye. Significant prognostic factors include geometric characteristics of the vessels – such as diameter, length and branching angles – as well as the distinction between arteries and veins, and their mutual arrangements in the eye. Those features are visible by fundus imaging (see example in figure 1), using a contrast agent to enhance the contrast of the vessels against their background. While manual examination of the progression of the disease is time consuming, ofttimes requiring integration and comparison of information from several modalities, algorithmic means emerge as a time-saving, fast and accurate means of detecting and tracking pathologies characterized by abnormalities in the vascular tree.
Segmentation of the vascular tree forms the first step towards the construction of an algorithmic basis for pathological detection. For this end, several binary classifiers have been proposed in the literature. One of the most recent, by Melinscak et al. of the University of Zagreb, offers a DNN approach for the construction of the binary classifier. In essence, for each pixel in a given image, the classifier relates it to a class – a vessel or not a vessel pixel (see the example of manual segmentation in figure 2, used for training of DNN). The choice of DNN seems a natural approach in this context, for which heterogeneity in the roles of processing layers in the network is required for recognizing different geometrical and spatial features of vessels, e.g. orientation of edges vs. color features. Melinscak’s 10 layered DNN underwent training in some of its layers, which included the filtering of the image between layers and the data propagated between them. It was then put to the test. By measuring the area under the Receiver Operating Characteristic (ROC), the researchers report an average accuracy of 0.96 to 0.98 in several experiments, which demonstrates the high potential and usability of DNN for automatic vessel segmentation.
The use of DNN and more specifically Convolution Neural Networks (CNN) for vessel segmentation, was demonstrated by Wang at el. In their approach, the CNN performs as a trainable hierarchical feature extractor. Combined with decision making based on random trees techniques, the network learns from intermediate layers as well as the output one.
Distinguishing veins from arteries forms the second step towards the detection of vascular tree pathologies. These two entities are oftentimes indistinguishable in imaging. Accurate diagnosis relies on the examiner to be able to tell apart vein and artery tree. For example, abnormal size of arteries is common in diabetes, whereas pancreas patients have narrower artery profiles. In this context, deep learning methodologies have also been demonstrated to perform well in automatic classification.
Relying on segmentation methods, whether based on DNNs or image processing techniques, the challenge is to train a binary classifier assigning each pixel as belonging to a vein or to an artery. Several algorithms exist to automatically perform this classification and most rely on just a few features distinguishing arteries from veins. Arteries are usually brighter and thinner than veins; the central reflex in arteries is wider than in veins; and near the optical disk veins and arteries alternate in position. Those features might prove insufficient in the case of lack of sufficient image contrast, or insufficient geometrical distinguishing feature near the outer parts of the image.
Automatic distinction between arteries and veins has been demonstrated using classification methods by Claudia Kondermann et al. of the University of Heidelberg. In their approach, after segmentation of the whole vascular tree was made, each pixel was assigned a tag of vein or artery in fundus images. Neural networks and Support Vector Machine were used to give an average accuracy of 95% in classification.
Taken together, the examples described above provide only the tip of the iceberg of the potential of DNN and neural networks for the classification and segmentation of vessels in fundus images. The advantage of learning algorithms are their flexibility and the relative ease of integrating new knowledge into the classification procedure. With the high percentage of classification accuracy, these algorithms can form the basis for the 3D reconstruction of the vascular tree, in which the distinction between veins and the artery tree can be made visible.References:
Distinguishing veins from arteries forms the second step towards the detection of vascular tree pathologies. These two entities are oftentimes indistinguishable in imaging. Accurate diagnosis relies on the examiner to be able to tell apart vein and artery tree. For example, abnormal size of arteries is common in diabetes, whereas pancreas patients have narrower artery profiles. In this context, deep learning methodologies have also been demonstrated to perform well in automatic classification.
Relying on segmentation methods, whether based on DNNs or image processing techniques, the challenge is to train a binary classifier assigning each pixel as belonging to a vein or to an artery. Several algorithms exist to automatically perform this classification and most rely on just a few features distinguishing arteries from veins. Arteries are usually brighter and thinner than veins; the central reflex in arteries is wider than in veins; and near the optical disk veins and arteries alternate in position. Those features might prove insufficient in the case of lack of sufficient image contrast, or insufficient geometrical distinguishing feature near the outer parts of the image.
Automatic distinction between arteries and veins has been demonstrated using classification methods by Claudia Kondermann et al. of the University of Heidelberg. In their approach, after segmentation of the whole vascular tree was made, each pixel was assigned a tag of vein or artery in fundus images. Neural networks and Support Vector Machine were used to give an average accuracy of 95% in classification.
Taken together, the examples described above provide only the tip of the iceberg of the potential of DNN and neural networks for the classification and segmentation of vessels in fundus images. The advantage of learning algorithms are their flexibility and the relative ease of integrating new knowledge into the classification procedure. With the high percentage of classification accuracy, these algorithms can form the basis for the 3D reconstruction of the vascular tree, in which the distinction between veins and the artery tree can be made visible.References:
- Lončarić, Sven, and Martina Melinščak. “Retinal Vessel Segmentation Using Deep Neural Networks.” VISAPP 2015 (10th International Conference on Computer Vision Theory and Applications). Hrvatska znanstvena bibliografija i MZOS-Svibor, 2015.
- Wang, et al., Hierarchical retinal blood vessel segmentation based on feature and ensemble learning, Neurocomputing (2014),
- Kondermann, Claudia, Daniel Kondermann, and Michelle Yan. “Blood vessel classification into arteries and veins in retinal images.” Medical Imaging. International Society for Optics and Photonics, 2007.

 Cardiology
Cardiology